|
 |
| Korean J Ophthalmol > Volume 36(5); 2022 > Article |
|
Abstract
MicroRNAs (miRNAs) are the small noncoding RNA molecules which regulate target gene expression posttranscriptionally. They are known to regulate key cellular processes like inflammation, cell differentiation, cell proliferation, and cell apoptosis across various ocular diseases. Due to their easier access, recent focus has been laid on the investigation of miRNA expression and their involvement in several conjunctival diseases. The aim of this narrative review is to provide understanding of the miRNAs and describe the current role of miRNAs as the mediators of the various conjunctival diseases. A literature search was made using PubMed, Scopus, and Web of Science databases for studies involving miRNAs in the conjunctival pathological conditions. Original articles in the last 10 years involving both human and animal models were included. Literature search retrieved 27 studies matching our criteria. Pertaining to the numerous literatures, there is a strong correlation between miRNA and the various pathological conditions that occur in the conjunctiva. miRNAs are involved in various physiological processes such as cell differentiation, proliferation, apoptosis, development, and inflammation by regulating various signaling pathways, genes, proteins, and mediators. Pterygium was the most studied conjunctival disease for miRNA involvement, whereas miRNA research in allergic conjunctivitis is still in its early stages. Our review provides deep insights into the various miRNAs playing an important role in the various conjunctival diseases. miRNAs do have the potential to serve as noninvasive biomarkers with diagnostic, prognostic, and therapeutic implications. However, multitudinous studies are required to validate miRNAs as the reliable biomarkers in conjunctival pathologies and its targeted therapy.
Since their discovery, knowledge about the pathophysiological role of microRNAs (miRNAs) in ocular surface has grown rapidly [1]. miRNAs are small noncoding RNA molecules of approximate 22 nucleotides in length that regulate gene expression posttranscriptionally [2]. They are known to regulate about one-third of human genome [3]. They negatively regulate the gene expression by targeting the 3′ untranslated region (UTR) region of target messenger RNA (mRNA). Hence, one miRNA can silence multiple target genes [4]. miRNAs are very resistant to degrading enzymes and are present across various body fluids such as tears, saliva, urine, plasma, and serum in a stable form. Thus, making them easily extracted and processed to serve as potential disease diagnostic and prognostic biomarkers [5].
In this review, we have consolidated various miRNA studies among conjunctival diseases from PubMed, Scopus, and Web of Science databases. We performed a literature search using the keywords “MicroRNAs”[MeSH] AND “Conjunctiva”[MeSH] OR “Conjunctival diseases”[ MeSH]. Our initial search retrieved a total of 51 articles. Following subsequent screening for articles solely dealing with the role of miRNAs in conjunctival tissue and conjunctival diseases, a total of 28 studies were extensively reviewed in this study. Relevant studies conducted both in human and animal models in the last 10 years, between January 2011 and December 2021, were included. Studies solely dealing with other epigenetic factors and other diseases were excluded. We aimed to comprehend and compare the studies carried out describing the expression and functions of various miRNAs in causing various conjunctival diseases.
In 1993, the first miRNA, lin-4, was discovered in Caenorhabditis elegans promoting the progression of initial larval stages by binding to the complementary sites of 3′ UTR mRNA region of lin-14 [1]. Seven years later, an associated gene called let-7 was discovered with a regulatory role of transmitting larval forms to an adult stage [6]. Since both let-4 and let-14 had common function of development process, they were referred to as “small temporal RNAs.” Their counterparts were reported the next year in humans and other species [7]. However, they were not present in all stages of development and were found only in specific cell types. Hence the term “microRNA” was presented to describe all these small RNA molecules with unknown functions. Subsequent studies suggested their regulatory role in gene expression posttranscriptionally by silencing their mRNA targets [8].
The miRNA biogenesis is a multiple step complex process starting from cytoplasm and ending in nucleus. A detailed canonical step-by-step pathway of miRNA biogenesis is shown in Fig. 1. In the first step, miRNA genes are transcribed by RNA polymerase II or RNA polymerase III into primary miRNA (pri-miRNA) transcripts [9]. The pri-miRNAs are generally over 1 kilobase long with one or more hairpin structures capped at 5′ and poly A tail [10]. In the next step, a nuclear ribonuclease III enzyme called Drosha, along with DiGeorge syndrome critical region 8, forms a microprocessor complex and cleaves the pri-miRNAs, giving out smaller precursors of miRNAs (pre-miRNAs) and initiating the maturation process [11]. These pre-miRNAs are then exported to nucleus from cytoplasm through exportin-5 and Ran-GTP (ras-related nuclear protein guanosine triphosphate) protein complex [12].
The pre-miRNAs in nucleus are further cleaved into 22-nucleotide-long miRNA duplexes by Dicer and transactivating response RNA-binding protein complex [13]. Now the functional mature strand in miRNA duplex is bounded to Argonaute proteins forming miRNA-induced silencing complex, which regulates the target gene expression through inhibiting translation or mRNA degradation. While the other passenger strand in miRNA duplex is degraded [14].
miRNAs can be easily accessed from various tissues and are also found across different body fluids such as plasma, serum, saliva, tears, blood, and urine [15]. To investigate conjunctival pathophysiology, miRNAs can be extracted from various sample sources, including conjunctival tissues or conjunctival cells, and tear fluid. To obtain conjunctival tissues, conjunctival cells, and tear fluid, various sampling methods such as surgical removal, impression cytology, and Schirmer’s test strips or capillary tubes are used. Generally, they are isolated from other molecules using various commercially available RNA extraction techniques. Different isolation techniques yield distinct concentrations of RNA. Thus, there is a need for an optimized extraction method based on the sample source and sample volume [16]. Qiagen miRNeasy Micro Kit (Qiagen, Hilden, Germany) is commonly used kit protocol designed to isolate miRNAs and small RNAs even from the small amounts of sample source. Bioanalyzer is an instrument that uses chip based microcapillary electrophoresis technique to assess the quality of total RNA and quantifying the small RNA and miRNA concentrations. Quantitative real-time polymerase chain reaction (RT-PCR), miRNA array analysis, and next generation sequencing are the most common methods to quantify and screen for active miRNAs from various body fluids. miRNAs may either be present as extracellular free circulating or enclosed in vesicles such as exosomes. Exosomes are generally considered to be rich source of miRNAs and many studies aim to extract exosomal miRNAs through ultracentrifugation for better yield [17].
Although it is known that the miRNAs play role in various biological processes such as cell growth, cell apoptosis, development, and differentiation across different tissues, their role in eye transcriptome is still unknown [18,19]. Defects in the function of the miRNAs are found to have intense effects on development [20,21].
Initial studies by Wienholds et al. [22] and Karali et al. [19] reported a spatiotemporal expression of 13 tissue specific miRNAs in the zebrafish eye and seven differentially expressed miRNAs in the mammalian eye, respectively. During early embryogenesis, the absence of miR-450b-5p and presence of miR-180 along with Pax6, a modulator of eye development, drives the differentiation and development of epithelia with ocular surface characteristics [23]. The miRNAs, miRNA-143/145 cluster and miR-145, were reported to affect epithelial cell processes such as proliferation and differentiation [24]. The miR-180 was found to be tissue specific and abundantly present in corneal tissue, indicating that it plays major role in maintaining healthy cornea [25]. Another study revealed increasing number of miRNAs being expressed at different embryonic developmental stages indicating their potential role in differentiation and development of progenitor cells [26]. However, further confirmatory and functional studies will elucidate their roles in developmental process of the eye.
Conjunctiva is a transparent mucus membrane lining the inside of the eyelids (palpebral conjunctiva) and sclera (bulbar conjunctival) [27]. In the recent years, a handful of studies on miRNA expression in the conjunctival tissues have revealed the involvement of miRNAs in the functions of the eyes. These studies have led to the further exploring the functions of miRNAs in conjunctiva and its related diseases. Altogether, disturbances on the miRNA expression may cause diseases responsible for conjunctival inflammation, infection, and blindness. The comprehensive relationships with the miRNAs are outlined in the Table 1 and is discussed ahead [28-54], while Fig. 2 summarizes the role of miRNAs in conjunctival disorders.
Allergic conjunctivitis (AC) is an inflammation of the conjunctiva in response to an allergen. It is one of the most common ocular conditions in the clinical practice. Though it is assumed that type 1 hypersensitivity, immunoglobulin E (IgE) and non-IgE-mediated mechanisms are responsible for its development, the pathophysiology of ocular allergy is complex and multifactorial, involving various mediators, chemokines, cytokines, receptors, and regulating pathways [55].
Sun et al. [28] designed an in vivo pollen induced mice model of AC and an in vitro lipopolysaccharide-stimulated inflammatory model of human corneal limbal epithelial cells. In these AC models, it was demonstrated that the eosinophils and total inflammatory cells were highly expressed and the expression of miR-146a was significantly reduced, while the expression of thymic stromal lymphopoietic protein (TSLP) and its downstream molecules was enhanced. Thus, it is reported that the downregulation of miR-146a induces AC by increasing TSLP levels [28]. A similar study 2 years later in a murine model, demonstrated that the miR-146a also contributes to the development of AC through regulating the inhibitory effect of regulatory T cells on conventional T cells a nd nuclear factor-κB (NF-κB) signaling pathway [29]. Moreover, another study in murine AC model, demonstrated the reduced levels of miR-19b while the levels of TSLP and phosphorylated signal transducer and activator of transcription 3 (STAT3) were increased suggesting that the miR-19b reduces the conjunctival inflammation through STAT3 signaling via TSLP downregulation [30]. Furthermore, a recent study elucidated the role and mode of action of miR-155 on AC in mice. The miR-155 was upregulated in the AC models while the inhibition of miR-155 reduces the AC-induced injury through a pathway inhibiting the phosphorylation of P65 [31].
These findings reveal the clinical potential of various miRNAs and their targets in the treatment of AC. The miRNA studies in AC are still confined to the murine models. Further studies in humans are needed to validate their potential in providing a basis for new drug research and development.
Numerous studies have reported dysregulation of miRNA expression in human cancers and abnormal expression of miRNAs is linked to cancer initiation and progression by functioning either as oncogenes or tumor suppressors [56,57]. Several miRNAs are reported to be differentially regulated across various lymphoma subtypes [58,59]. Conjunctival lymphoma is a malignant ocular surface tumor arising from B-cell lymphocytes [60]. It accounts for about 25% of ocular adnexal lymphomas (OALs) [61].
The miRNA microarray profiling of mucosal-associated lymphoid tissue (MALT) lymphoma, the most common subtype of OALs and normal adjacent tissues by Cai et al. [32] revealed dysregulation of miRNAs. The miR-150 and miR-155 were upregulated and the miR-184, miR-200a, b, c, and miR-205 were downregulated. It was further demonstrated that the dysregulation of miR-200 family could be linked in the pathogenesis of conjunctival MALT lymphoma [32]. A year later, another study investigated the expression of miRNAs between low-grade extranodal marginal zone lymphoma and diffuse large B-cell lymphoma, the two other major subtypes of OALs. The miRNA expression profiling revealed 43 differentially expressed miRNAs. The differential miRNA expression is probably due to the differences in MYC protein and NF-κB regulatory pathways [33].
Conjunctival melanoma (CM) is presented as the raised pigmented or nonpigmented lesion of the ocular surface [62]. Genetic studies revealed mutations in the somatic genes, BRAF, NRAS, NFI, KIT, and TERT, suggesting that CMs are more closely related to skin and mucosal melanomas [63,64]. Since none of these mutations can predict the outcome of the disease, it demands a need of novel molecular markers that can predict the clinical course of CM.
A distinct study in 2016 performed the microarray analysis between CMs and normal conjunctival samples revealing 24 upregulated and one downregulated miRNAs in conjunct [34]. Several miRNAs were associated with the tumor thickness and specific for stage-T1 and stage-T2 of CM, while miR-3687 and miR-3916 were associated with an increased risk of regional recurrence. The study further highlighted the resemblance between CM, mucosal melanoma, and cutaneous melanoma [34]. Another recent study investigated the miRNA profile in primary tumors from nonmetastasizing and metastasizing CM. Two different sets of miRNAs were differentially expressed in two CM forms suggesting that these miRNAs can predict the metastatic progression. The miR-184 was downregulated in the progressive stages of CM, seemingly involved in driving the tumor cells into the more severe metastatic form [35]. These profiling studies confirm the role of miRNAs in CM and provide an entry point for future functional studies of miRNAs as prognostic or therapeutic targets in CM.
Numerous studies have demonstrated the role of miRNAs in the differentiation across various cell types. For example, the role of miR-150 in B cell differentiation and miR-196a in osteogenic differentiation [65,66]. Only recently the role of miRNAs in conjunctival epithelial cell differentiation has been explored.
Ranjbarnejad et al. [36] explored the role of miR-let-7a on conjunctival mesenchymal stem cells differentiation into photoreceptor-like cells. It has been previously demonstrated that the upregulation of miR-let-7a accelerates the development in the retina [36]. While investigating its role in conjunctival mesenchymal stem cell differentiation, it is shown that the overexpression of miR-let-7a increases the expression of photoreceptor-specific genes, and its effect is time dependent [67]. Another recent study has investigated the role of miRNAs in modulating the epithelial to mesenchymal transitions (EMT) in human conjunctival epithelial cells. The RT-PCR-based array directed at members of miR-200 family as the key regulators of EMT process in human conjunctival epithelial cells. It further indicated that DNA demethylation of promoters of miR-200 loci is very important for reverting the EMT in human conjunctival epithelial cells, suggesting the prospective for the developing new epigenetic-based therapeutic methods for treating conjunctival conditions associated with EMT [37].
Pterygium is a degenerative ocular surface disease where an abnormal ingrowth of conjunctival tissue pervades the peripheral cornea. The pathogenesis of pterygium is a complex process involving inflammation, neovascularization, abnormal proliferation, and apoptosis. It is generally associated with the long-term ultraviolent radiation exposure [68,69]. Since it is known that miRNAs are the key regulators of differentiation, EMT processing and apoptosis in various conditions and their regulatory role in the primary pterygium is investigated by a handful of studies.
The correlation between miR-145 expression levels and the clinical severity of pterygium was investigated in 253 patients using RT-PCR by Chien et al. [38]. They reported a negative correlation between miR-145 expression and pterygium characteristics suggesting more studies on the miR-145 as a potential treatment due to its known tumor suppression effect [38]. Another study carried out microarray profiling to determine the differentially expressed miRNAs in pterygium compared to normal conjunctiva. There were 25 differentially expressed miRNAs in total: 14 upregulated and 11 downregulated. The collective downregulation of the miR-200 family, known to regulate the EMT, suggests that EMT could potentially play role in the pathogenesis of pterygium [39,69]. Subsequently Wu et al. [40] has reiterated the role of miR-200a in EMT processing and pterygium pathogenesis.
β-catenin is a multifunctional protein that contributes to cell development and is said to involve in pterygium pathogenesis along with EMT [70]. Wu et al. [41] investigated the hypothesis, that the β-catenin, miR-221, and its downstream p27Kipl gene expression were correlated with the pathogenesis of pterygium. Most of the pterygium specimens (60%) had high expression levels of miR-221 compared to the control groups. Furthermore, miR-221 negatively regulated the p27Kipl gene expression in pterygium. Thus, upon activation, the β-catenin interacts with miR-221, resulting in the downregulation of the p27Kipl gene in the pathogenesis of pterygium [41]. In the ocular surface pterygium fibroblast cells, the miR-215 could take part in inhibiting the fibroblast proliferation while in the human pterygium epithelial cells, the miR-218-5p is said to inhibit the migration and proliferation of the human pterygium epithelial cells [42,43]. Investigating the role of apoptosis in the pathogenesis of pterygium, the miR-122 is reported to inversely regulate the expression of Bcl-w, an antiapoptosis protein, resulting in abnormal cell apoptosis leading to the development of pterygium [44].
The expression of miR-21, a cancer-promoting miRNA in humans, is upregulated and is increased as the severity of the pterygium increased. Thereby, miR-21 promotes pterygium cell differentiation through PTEN/AKT pathway, and its inhibition suppressed the proliferation and induces apoptosis of pterygium fibroblast cells [45]. The miR-143 and miR-145 are predominantly expressed in the basal epithelia of pterygium [46]. The miR-145 reduced the expression of an oncogene, MDM2 which subsequently affected the p53-related growth pattern [46]. Meanwhile, another miRNA, miR-3175 is shown to promote proliferation, migration, invasion, and EMT in human conjunctiva and pterygium by directly inhibiting Smad7 [47]. Additionally, a recent study reported the expression levels of miR-182-5p, miR-183-5p, and miR-184 to be increased while the expression of miR-221-3p was decreased in the primary pterygium in comparison with the normal conjunctiva [48]. The expression of miR-143/145 mediates TGF-β-induced human subconjunctival fibrosis [49].
Steven-Johnson syndrome (SJS) is a rare, serious inflammatory vesiculobullous reaction of skin and mucosa such as oral cavity, ocular surface, and genitals. The progressive forms are called toxic epidermal necrolysis (TEN) [71]. About half of the SJS/TEN patients show severe ocular complications (SOC). Acute cases of SJS/TEN with SOC will present severe conjunctivitis with corneal and conjunctival erosion while the chronic cases will have persistent conjunctival inflammation [72]. The exact pathophysiology of SJS is not fully understood. Delayed drug hypersensitivity reactions to a drug or drug metabolite are the commonly attributed reason in most cases. Only a handful of studies have illustrated the role of miRNAs in TEN. The expression of miR-18a-5p, miR-124, and miR-375-3p were upregulated in various cells and body fluids [73-75]. Ueta et al. [50] investigated the role of miRNAs in the pathogenesis of SJS/TEN patients with SOC. The expression of miR-31 and miR-455-3p is shown to be significantly upregulated in the conjunctival epithelium of SJS/ TEN chronic patients. It was further reported that the miR-455-3p regulates many innate immune related genes in human conjunctival epithelium [50].
Trachoma is a contagious bacterial infection caused by the bacterium Chlamydia trachomatis that infects the eyes. It is a leading infectious disease that causes blindness [76]. The initial symptoms include mild itching and irritation of the eyes slowly progressing to swelling and pus discharge. Persistent forms of trachomatous is believed to drive the continued scarring process even in the absence of C. trachomatis infection [77]. The miRNA expression is known to be dysregulated upon bacterial infection [78].
Derrick et al. [51] investigated the expression of miRNAs in normal and trachomatous conjunctival swaps. The expression of miR-147b and miR-1285 were reported to be upregulated in the inflammatory trachomatous scarring [51]. The same team later investigated the miRNAs expression during the initial stage of disease (follicular trachoma) with active infection. The quantitative PCR data validated the expression of nine miRNAs, miR-155, miR-150, miR-142, miR-181b, miR-181a, miR-132, miR-342, miR-184, and miR-4728, to be differentially expressed during follicular trachoma. In particular, the expression of miR-155 (upregulated) and miR-184 (downregulated) is correlated with the severity of inflammation [52]. Now that these miRNAs were associated with the trachomatous inflammation and infections, their role in predicting the disease progression was investigated. Conjunctival swabs of 506 samples served as the baseline sample set for 4-year longitudinal study. However, none of the miRNAs, miR-147b, miR-1285, miR-184, and miR-155, were associated with 4-year scarring incidence and progression [53]. Additionally, a recent study revealed 21 differentially expressed miRNAs in a cohort of patients with progressive limbal stem cell deficiency. Of these, miR-204-5p, an inhibitor of corneal neovascularization, was significantly downregulated [54].
In conclusion, this review summarizes the biogenesis and differential expression of various miRNAs in numerous conjunctival pathologies. miRNAs play a key role in different cell processes such as cell proliferation, inflammation, differentiation, and cell development. Although miRNAs have been extensively studied across various diseases since their discovery over two decades ago, miRNA research in ocular diseases is still in its infancy. Our study faced limitations in reviewing due to the scarcity of literature in eye diseases, especially conjunctival diseases. Furthermore, we felt that there was a need for an optimizing study for different miRNA isolation methods from different samples of conjunctiva. miRNAs have emerged as the key regulators of inflammation in allergic diseases. Despite the fact that the majority of patients with allergic conjunctivitis have a familial history of atopy and a genetic predisposition to develop the disease, it remains the least explored conjunctival disease. Moreover, there were no reported studies exploring the role of miRNAs in more chronic and recurring conjunctival diseases such as vernal keratoconjunctivitis. Much of the miRNA research in conjunctival diseases are focused on pterygium and ocular cancers, owing to the extensive literature available on other systemic diseases such as cancers.
Several studies highlight the need to integrate miRNA expression studies on biological processes in order to fully understand eye diseases. Current advancements in technology will enable us to develop better diagnostic and therapeutic strategies for conjunctival diseases. The miRNAs can serve as noninvasive potential biomarkers with diagnostic, prognostic, and therapeutic implications. However, further research is needed to assert whether regulating the miRNAs associated with the diseases can control the diseases.
References
1. Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993;75:843-54.


2. Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 2010;11:597-610.


4. Ye J, Xu M, Tian X, et al. Research advances in the detection of miRNA. J Pharm Anal 2019;9:217-26.



6. Reinhart BJ, Slack FJ, Basson M, et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000;403:901-6.


8. Rassi DM, De Paiva CS, Dias LC, et al. Review: microRNAs in ocular surface and dry eye diseases. Ocul Surf 2017;15:660-9.


9. O’Brien J, Hayder H, Zayed Y, Peng C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne) 2018;9:402.


11. Achkar NP, Cambiagno DA, Manavella PA. miRNA biogenesis: a dynamic pathway. Trends Plant Sci 2016;21:1034-44.


12. Kawahara Y. Human diseases caused by germline and somatic abnormalities in microRNA and microRNA-related genes. Congenit Anom (Kyoto) 2014;54:12-21.


13. Qiu X, Zhang H, Yu H, et al. Duplex-specific nuclease-mediated bioanalysis. Trends Biotechnol 2015;33:180-8.


14. Lao TD, Le TA. MicroRNAs: biogenesis, functions and potential biomarkers for early screening, prognosis and therapeutic molecular monitoring of nasopharyngeal carcinoma. Processes 2020;8:966.

15. Weber JA, Baxter DH, Zhang S, et al. The microRNA spectrum in 12 body fluids. Clin Chem 2010;56:1733-41.



16. Brown RA, Epis MR, Horsham JL, et al. Total RNA extraction from tissues for microRNA and target gene expression analysis: not all kits are created equal. BMC Biotechnol 2018;18:16.



17. Nik Mohamed Kamal NN, Awang RA, Mohamad S, Shahidan WN. Plasma- and saliva exosome profile reveals a distinct microRNA signature in chronic periodontitis. Front Physiol 2020;11:587381.



18. Macfarlane LA, Murphy PR. MicroRNA: biogenesis, function and role in cancer. Curr Genomics 2010;11:537-61.



19. Karali M, Peluso I, Marigo V, Banfi S. Identification and characterization of microRNAs expressed in the mouse eye. Invest Ophthalmol Vis Sci 2007;48:509-15.


20. Wienholds E, Koudijs MJ, van Eeden FJ, et al. The microRNA-producing enzyme Dicer1 is essential for zebrafish development. Nat Genet 2003;35:217-8.


21. Bernstein E, Kim SY, Carmell MA, et al. Dicer is essential for mouse development. Nat Genet 2003;35:215-7.


22. Wienholds E, Kloosterman WP, Miska E, et al. MicroRNA expression in zebrafish embryonic development. Science 2005;309:310-1.


23. Shalom-Feuerstein R, Serror L, De La Forest Divonne S, et al. Pluripotent stem cell model reveals essential roles for miR-450b5pand miR-184 in embryonic corneal lineage specification. Stem Cells 2012;30:898-909.


24. Lee ST, Chu K, Im WS, et al. Altered microRNA regulation in Huntington’s disease models. Exp Neurol 2011;227:172-9.


25. Abu-Amero KK, Helwa I, Al-Muammar A, et al. Screening of the seed region of MIR184 in keratoconus patients from Saudi Arabia. Biomed Res Int 2015;2015:604508.



26. Xu S. microRNA expression in the eyes and their significance in relation to functions. Prog Retin Eye Res 2009;28:87-116.


27. Yanoff M, Sassani JW. Ocular pathology. 7th ed. Philadelphia: Saunders; 2014.
28. Sun W, Sheng Y, Chen J, et al. Down-regulation of miR-146a expression induces allergic conjunctivitis in mice by increasing TSLP level. Med Sci Monit 2015;21:2000-7.



29. Yang Y, Yin X, Yi J, Peng X. MiR-146a overexpression effectively improves experimental allergic conjunctivitis through regulating CD4+CD25-T cells. Biomed Pharmacother 2017;94:937-43.


30. Guo C, Liu J, Hao P, et al. The potential inhibitory effects of miR-19b on ocular inflammation are mediated upstream of the JAK/STAT pathway in a murine model of allergic conjunctivitis. Invest Ophthalmol Vis Sci 2020;61:8.



31. Bai M, Li Y, Li Y, Hu Z. Influence of miR155 on allergic conjunctivitis in mice via regulation of NF-κB signal pathway. Trop J Pharm Res 2019;18:1603-8.
32. Cai J, Liu X, Cheng J, et al. MicroRNA-200 is commonly repressed in conjunctival MALT lymphoma, and targets cyclin E2. Graefes Arch Clin Exp Ophthalmol 2012;250:523-31.


33. Hother C, Rasmussen PK, Joshi T, et al. MicroRNA profiling in ocular adnexal lymphoma: a role for MYC and NFKB1 mediated dysregulation of microRNA expression in aggressive disease. Invest Ophthalmol Vis Sci 2013;54:5169-75.


34. Larsen AC, Mikkelsen LH, Borup R, et al. MicroRNA expression profile in conjunctival melanoma. Invest Ophthalmol Vis Sci 2016;57:4205-12.


35. Mikkelsen LH, Andersen MK, Andreasen S, et al. Global microRNA profiling of metastatic conjunctival melanoma. Melanoma Res 2019;29:465-73.


36. Ranjbarnejad F, Nadri S, Biglari A, et al. Effect of let-7a overexpression on the differentiation of conjunctiva mesenchymal stem cells into photoreceptor-like cells. Iran J Basic Med Sci 2019;22:878-83.


37. Rajic J, Dinic S, Uskokovic A, et al. DNA methylation of miR-200 clusters promotes epithelial to mesenchymal transition in human conjunctival epithelial cells. Exp Eye Res 2020;197:108047.


38. Chien KH, Chen SJ, Liu JH, et al. Correlation of microRNA-145 levels and clinical severity of pterygia. Ocul Surf 2013;11:133-8.


39. Engelsvold DH, Utheim TP, Olstad OK, et al. miRNA and mRNA expression profiling identifies members of the miR-200 family as potential regulators of epithelial-mesenchymal transition in pterygium. Exp Eye Res 2013;115:189-98.



40. Wu CW, Peng ML, Yeh KT, et al. Inactivation of p53 in pterygium influence miR-200a expression resulting in ZEB1/ZEB2 up-regulation and EMT processing. Exp Eye Res 2016;146:206-11.


41. Wu CW, Cheng YW, Hsu NY, et al. MiRNA-221 negatively regulated downstream p27Kip1 gene expression involvement in pterygium pathogenesis. Mol Vis 2014;20:1048-56.


42. Lan W, Chen S, Tong L. MicroRNA-215 regulates fibroblast function: insights from a human fibrotic disease. Cell Cycle 2015;14:1973-84.



43. Han S, Chen Y, Gao Y, et al. MicroRNA-218-5p inhibit the migration and proliferation of pterygium epithelial cells by targeting EGFR via PI3K/Akt/mTOR signaling pathway. Exp Eye Res 2019;178:37-45.


44. Cui YH, Li HY, Gao ZX, et al. Regulation of apoptosis by miR-122 in pterygium via targeting Bcl-w. Invest Ophthalmol Vis Sci 2016;57:3723-30.


45. Li X, Dai Y, Xu J. MiR-21 promotes pterygium cell proliferation through the PTEN/AKT pathway. Mol Vis 2018;24:485-94.


46. Teng Y, Yam GH, Li N, et al. MicroRNA regulation of MDM2-p53 loop in pterygium. Exp Eye Res 2018;169:149-56.


47. Zhong X, Tang J, Li H, et al. MiR-3175 promotes epithelial-mesenchymal transition by targeting Smad7 in human conjunctiva and pterygium. FEBS Lett 2020;594:1207-17.


48. Icme G, Yilmaz A, Dinc E, et al. Assessment of miR-182, miR-183, miR-184, and miR-221 expressions in primary pterygium and comparison with the normal conjunctiva. Eye Contact Lens 2019;45:208-11.


49. Hwang YH, Jung SA, Lyu J, et al. Transforming growth factor-β1-induced human subconjunctival fibrosis is mediated by microRNA 143/145 expression. Invest Ophthalmol Vis Sci 2019;60:2064-71.


50. Ueta M, Nishigaki H, Sotozono C, et al. Regulation of gene expression by miRNA-455-3p upregulated in the conjunctival epithelium of patients with Stevens-Johnson syndrome in the chronic stage. Sci Rep 2020;10:17239.



51. Derrick T, Roberts Ch, Rajasekhar M, et al. Conjunctival MicroRNA expression in inflammatory trachomatous scarring. PLoS Negl Trop Dis 2013;7:e2117.



52. Derrick T, Last AR, Burr SE, et al. Inverse relationship between microRNA-155 and −184 expression with increasing conjunctival inflammation during ocular Chlamydia trachomatis infection. BMC Infect Dis 2016;16:60.



53. Derrick T, Ramadhani AM, Mtengai K, et al. miRNAs that associate with conjunctival inflammation and ocular Chlamydia trachomatis infection do not predict progressive disease. Pathog Dis 2017;75:ftx016.



54. Latta L, Ludwig N, Krammes L, et al. Abnormal neovascular and proliferative conjunctival phenotype in limbal stem cell deficiency is associated with altered microRNA and gene expression modulated by PAX6 mutational status in congenital aniridia. Ocul Surf 2021;19:115-27.


55. La Rosa M, Lionetti E, Reibaldi M, et al. Allergic conjunctivitis: a comprehensive review of the literature. Ital J Pediatr 2013;39:18.



56. Peng Y, Croce CM. The role of MicroRNAs in human cancer. Signal Transduct Target Ther 2016;1:15004.



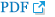
57. Negrini M, Nicoloso MS, Calin GA. MicroRNAs and cancer: new paradigms in molecular oncology. Curr Opin Cell Biol 2009;21:470-9.


58. Lawrie CH, Soneji S, Marafioti T, et al. MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma. Int J Cancer 2007;121:1156-61.


59. Eis PS, Tam W, Sun L, et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci U S A 2005;102:3627-32.



60. Tanenbaum RE, Galor A, Dubovy SR, Karp CL. Classification, diagnosis, and management of conjunctival lymphoma. Eye Vis (Lond) 2019;6:22.



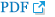
61. Dalvin LA, Salomao DR, Patel SV. Population-based incidence of conjunctival tumours in Olmsted County, Minnesota. Br J Ophthalmol 2018;102:1728-34.



62. Wong JR, Nanji AA, Galor A, Karp CL. Management of conjunctival malignant melanoma: a review and update. Expert Rev Ophthalmol 2014;9:185-204.



63. Mikkelsen LH, Larsen AC, von Buchwald C, et al. Mucosal malignant melanoma: a clinical, oncological, pathological and genetic survey. APMIS 2016;124:475-86.


64. Larsen AC, Dahl C, Dahmcke CM, et al. BRAF mutations in conjunctival melanoma: investigation of incidence, clinicopathological features, prognosis and paired premalignant lesions. Acta Ophthalmol 2016;94:463-70.


65. Xiao C, Calado DP, Galler G, et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 2007;131:146-59.


66. Kim YJ, Bae SW, Yu SS, et al. miR-196a regulates proliferation and osteogenic differentiation in mesenchymal stem cells derived from human adipose tissue. J Bone Miner Res 2009;24:816-25.


67. La Torre A, Georgi S, Reh TA. Conserved microRNA pathway regulates developmental timing of retinal neurogenesis. Proc Natl Acad Sci U S A 2013;110:E2362-70.


68. Chui J, Di Girolamo N, Wakefield D, Coroneo MT. The pathogenesis of pterygium: current concepts and their therapeutic implications. Ocul Surf 2008;6:24-43.


69. Ando R, Kase S, Ohashi T, et al. Tissue factor expression in human pterygium. Mol Vis 2011;17:63-9.


70. Kato N, Shimmura S, Kawakita T, et al. Beta-catenin activation and epithelial-mesenchymal transition in the pathogenesis of pterygium. Invest Ophthalmol Vis Sci 2007;48:1511-7.


71. Sotozono C, Ueta M, Nakatani E, et al. Predictive factors associated with acute ocular involvement in Stevens-Johnson syndrome and toxic epidermal necrolysis. Am J Ophthalmol 2015;160:228-37.


72. Ueta M, Nishigaki H, Sotozono C, Kinoshita S. Downregulation of interferon-γ-induced protein 10 in the tears of patients with Stevens-Johnson syndrome with severe ocular complications in the chronic stage. BMJ Open Ophthalmol 2017;1:e000073..



73. Sato S, Ichihara A, Jinnin M, et al. Serum miR-124 up-regulation as a disease marker of toxic epidermal necrolysis. Eur J Dermatol 2015;25:457-62.


74. Ichihara A, Wang Z, Jinnin M, et al. Upregulation of miR-18a-5p contributes to epidermal necrolysis in severe drug eruptions. J Allergy Clin Immunol 2014;133:1065-74.


75. Zhang C, Zhu Z, Gao J, et al. Plasma exosomal miR-375-3p regulates mitochondria-dependent keratinocyte apoptosis by targeting XIAP in severe drug-induced skin reactions. Sci Transl Med 2020;12:eaaw6142.


76. Mariotti SP, Pascolini D, Rose-Nussbaumer J. Trachoma: global magnitude of a preventable cause of blindness. Br J Ophthalmol 2009;93:563-8.


Fig. 1
Illustration of multistep canonical microRNA (miRNA) biogenesis pathway in humans. Pol = polymerase; pri-miRNA = primary miRNA; DGCR8 = DiGeorge syndrome critical region 8; pre-miRNA = precursors of miRNA; Ran = ras-related nuclear protein; GTP = guanosine triphosphate; TRBP = HIV-transactivating response RNA-binding protein; AGO = Argonaute; miRISC = miRNA-induced silencing complex; mRNA = messenger RNA. Created using BioRender (Toronto, Canada; https://biorender.com/).
Fig. 2
Summarized roles of the microRNAs across several conjunctival diseases. Created using BioRender (Toronto, Canada; https://biorender.com/).
Table 1
Differentially expressed miRNAs across various conjunctival disorders
| Study | Disease/condition | miRNAs | Expression | Functional role | Species |
|---|---|---|---|---|---|
| Sun et al. [28] | AC | miR-146a | Downregulation | Induces AC by increasing TSLP levels | Mice |
| Yang et al. [29] | AC | miR-146a | Upregulation | Regulates the inhibitory effect of Treg on Tcons and NF-κB pathway | Mice |
| Guo et al. [30] | AC | miR-19b | Downregulation | Reduction in allergic inflammation involving STAT3 signaling pathway | Mice |
| Latta et al. [31] | AC | miR-155 | Upregulation | Phosphorylation of P65 and increased inflammatory mediators | Mice |
| Cai et al. [32] | Conjunctival lymphoma | miR-200 family | Downregulation | Increased expression of cyclin E2 in diseased cells | Human |
| Hother et al. [33] | Conjunctival lymphoma | 43 miRNAs | Differential expression | Inhibition of MYC oncoprotein and NF-κB in aggressive forms | Human |
| Larson et al. [34] | CM | 25 miRNAs | Differential expression | Changes associated with tissue thickness and risk of relapse | Human |
| Mikkelsen et al. [35] | CM | 15 miRNAs | Differential expression | Prediction of metastatic phenotype by hsa-miR-184 | Human |
| Ranjbarnejad et al. [36] | Conjunctival cell differentiation | let-7a | Upregulation | Differentiation of conjunctival mesenchymal stem cells into photoreceptor-like cells | Human |
| Rajic et al. [37] | Conjunctival cell differentiation | miR-200 family | Downregulation | DNA methylation of promoters of miR-200 promotes EMT in human conjunctival epithelial cells | Human |
| Chien et al. [38] | Pterygium | miR-145 | Downregulation | Indirect regulation of angiogenesis and pterygia | Human |
| Engelsvold et al. [39] | Pterygium | miR-200 family | Downregulation | Induction of mesenchymal differentiation, and activation of transcription factors | Human |
| Wu et al. [40] | Pterygium | miR-200a | Downregulation | Increase in ZEB1/ZEB2 protein levels affecting EMT | Human |
| Wu et al. [41] | Pterygium | miR-221 | Upregulation | Downregulation of β-catenin associated p27Kipl gene | Human |
| Lan et al. [42] | Pterygium | miR-215 | Downregulation | Appearance of cell cycle related markers at G1/S and G2/M phases | Human |
| Han et al. [43] | Pterygium | miR-218-5p | Downregulation | Inhibiting proliferation of pterygium epithelial cells by targeting EGFR via PI3K/mTOR signaling pathway | Human |
| Cui et al. [44] | Pterygium | miR-122 | Downregulation | Regulates cell apoptosis via its regulation of Bcl-w expression | Human |
| Li et al. [45] | Pterygium | miR-21 | Upregulation | Inhibition of the PTEN/AKT axis and aberrant cell proliferation | Human |
| Teng et al. [46] | Pterygium | miR-145 | Upregulation | p53 associated patterned growth on attenuation of MDM2 | Human |
| Hwang et al. [47] | Pterygium | miR-3175 | Upregulation | Increase in EMT and migration with inhibition of Smad7 | Human |
| Icme et al. [48] | Pterygium | miR-182, miR-183, miR-184, miR-221 | Upregulation*, Downregulation† | Involvement of inflammatory and EMT pathway in pterygium | Human |
| Icme et al. [49] | Conjunctival fibrosis | miR-143, miR-145 | Downregulation | Mediates TGF-β induced human subconjunctival fibrosis | Human |
| Ueta et al. [50] | Stevens-Johnson syndrome | miR-455 | Upregulation | Increased activation of innate immune genes and cytokines | Human |
| Derrick et al. [51] | Trachomatis | miR-147b, miR-1285 | Upregulation | Initiates cell differentiation, inflammation | Human |
| Derrick et al. [52] | Trachomatis | miR-155, miR-184 | Upregulation‡, Downregulation§ | miRNA expression is a direct marker of inflammation | Human |
| Derrick et al. [53] | Trachomatis | miR-147b, miR-1285, miR-155, miR-184 | No significant expression | Do not reliably predict progression of scarring in trachoma patients | Human |
| Latta et al. [54] | Limbal stem cell deficiency | miR-204-5p | Downregulation | IEG activation and cell transduction leading to neovascularization | Human |
miRNA = microRNA; AC = allergic conjunctivitis; TSLP = thymic stromal lymphopoietic protein; Treg = regulatory T cell; Tcon = conventional T cell; NF-κB = nuclear factor-κB; STAT3 = signal transducer and activator of transcription 3; CM = conjunctival melanoma; EMT = epithelial to mesenchymal transitions; G1 = gap 1; S = synthesis; G2 = gap 2; M = mitosis; EGFR = epidermal growth factor receptor; P13K = phosphatidylinositol,3,4,5 triphosphate; mTOR = mammalian target of rapamycin; MDM2 = mouse double minute 2; TGF-β = transforming growth factor-β; IEG = immediate early gene.
- TOOLS



 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print